|
 |
| Tuberc Respir Dis > Volume 75(5); 2013 > Article |
|
Abstract
The emergence of new therapeutic agents for non-small cell lung cancer (NSCLC) implies that histologic subtyping and molecular predictive testing are now essential for therapeutic decisions. Histologic subtype predicts the efficacy and toxicity of some treatment agents, as do genetic alterations, which can be important predictive factors in treatment selection. Molecular markers, such as epidermal growth factor receptor (EGFR) mutation and anaplastic lymphoma kinase (ALK) rearrangement, are the best predictors of response to specific tyrosine kinase inhibitor treatment agents. As the majority of patients with NSCLC present with unresectable disease, it is therefore crucial to optimize the use of tissue samples for diagnostic and predictive examinations, particularly for small biopsy and cytology specimens. Therefore, each institution needs to develop a diagnostic approach requiring close communication between the pulmonologist, radiologist, pathologist, and oncologist in order to preserve sufficient biopsy materials for molecular analysis as well as to ensure rapid diagnosis. Currently, personalized medicine in NSCLC is based on the histologic subtype and molecular status. This review summarizes strategies for tissue acquisition, histologic subtyping and molecular analysis for predictive testing in NSCLC.
Until recently, the classification of lung cancer into two groups, non-small cell lung cancer (NSCLC) and small cell lung cancer, was adequate for selection of appropriate therapy. However, the use of new therapeutic agents for NSCLC requires definite knowledge of histological subtype. In particular, it is important to discriminate between squamous cell carcinoma and non-squamous cell carcinoma in NSCLC. In recent phase III clinical trials, pemetrexed showed a superior effect in non-squamous cell carcinoma, such as adenocarcinoma and large cell carcinoma, than in squamous cell carcinoma1,2. Bevacizumab is not recommended for treatment of squamous cell carcinoma, due to the higher frequency of pulmonary hemorrhage in this histologic subtype3,4.
The epidermal growth factor receptor-tyrosine kinase inhibitors (EGFR-TKIs) gefitinib and erlotinib produce a dramatic response when used as a first-line therapy in NSCLCs carrying an activating EGFR mutation5,6. The anaplastic lymphoma kinase (ALK) inhibitor, crizotinib, is effective in NSCLCs with rearrangement of ALK7. Moreover, the molecular markers for targeting of these agents, EGFR mutations and ALK rearrangements, are primarily found in adenocarcinoma5-7. Therefore, the histologic subtype of NSCLC is an important consideration, not only in selecting cytotoxic chemotherapeutic agents, but also to target molecular analysis tests.
Approximately 70% of patients with lung cancer are diagnosed using small biopsy or cytology specimens, as they present with unresectable, advanced stage disease8,9. Thus, it is important to be able to diagnose lung cancer and confirm histologic subtypes using small specimens.
While the amount of specimen required for histologic subtyping and molecular predictive testing has increased, the sample size obtained from patients is frequently insufficient. Therefore, it is critical to optimize the use of tissue samples for diagnostic and predictive examination10,11.
This article will review strategies for tissue acquisition, pathology approach, histologic subtyping, specimen requirement, and molecular analysis for predictive testing in NSCLC.
For a definite diagnosis of lung cancer, it is important to obtain adequate tissue samples and to process these adequately. Minimally invasive techniques can be used to obtain specimens in patients with advanced NSCLC. The available methods for acquisition of tumor cells or tissue are exfoliation cytology, aspiration cytology, and biopsy11.
Exfoliation cytology sampling techniques include bronchial washing, bronchoalveolar lavage, bronchial brushing, and sputum cytology. Aspiration cytology sampling methods include transbronchial needle aspiration (TBNA), endobronchial ultrasound (EBUS), and endoscopic ultrasound; these may be used in conjunction with one or more other techniques. Biopsies used to diagnose lung cancer include bronchial biopsy, transbronchial lung biopsy, and transthoracic needle biopsy11-13.
The success rates for these procedures vary considerably. The size of needle and forceps and the number of biopsies may affect the accuracy of diagnosis and risk of complications11,14.
Rapid on site examination of TBNA specimens is a fast cytological examination of sample adequacy and allows preliminary diagnosis by a pathologist11,12. In addition, the telepathology system makes it possible for pathologists to view cytology samples remotely and is a valuable time-saving technology12,15.
Cytology samples obtained by EBUS-TBNA are sufficient for evaluation of EGFR mutation and ALK rearrangement by molecular testing10,16,17.
The pulmonologist or radiologist who performs a tissue sampling procedure should endeavor to obtain sufficient tissue for diagnosis and molecular analysis. If only limited tissue is acquired, owing to unanticipated technical problems or complications during the procedure, the order of preference for tissue analysis must be considered8,9,12,18.
Accurate and relevant clinical information is essential for appropriate handling of small samples from suspected lung cancers in pathology laboratories. Five features providing clinical information relevant to pathology diagnosis are sample site, primary tumor or metastasis, presumptive diagnosis, previous relevant treatment (including surgery, chemotherapy, or radiotherapy), and smoking history11. Using this information, pathologists must determine priorities for the diagnostic approach and arrange necessary investigations.
Generally, the diagnostic approach to lung cancer workup consists of three steps: 1) identification of malignancy, 2) histologic typing, and 3) molecular analysis as appropriate11.
Reflex testing, a testing policy that does not require a clinician order for each case, may help to assure rapid diagnosis without delay of immunohistochemistry (IHC) and molecular testing. Conversely, this approach may not be optimal if tissue samples are insufficient10,11,18.
In about two thirds of NSCLC cases it is possible to differentiate between squamous cell carcinoma and adenocarcinoma adequately using morphological characteristics. In these cases, immunohistochemial staining is not required and small tissue samples can be preserved for appropriate molecular testing8-11,18.
The new classification of lung adenocarcinoma recommends that NSCLC is further identified as a specific histologic type, such as adenocarcinoma or squamous cell carcinoma, whenever possible8. Adenocarcinoma may manifest glandular differentiation with lepidic (formerly bronchioloalveolar), acinar, papillary, micropapillary, or solid growth patterns, while key morphologic features of squamous cell carcinoma are keratinization, pearl formation, and/or intercelluar bridges9.
The percentage of NSCLC cases that cannot be accurately classified into histologic subtypes can be as high as 40%; such cases are diagnosed as NSCLC-not otherwise specified (NOS)8,9,11. Recently, it was recommended that the diagnosis of NSCLC-NOS should be reduced to less than 5%, and IHC may be a useful method for refining diagnosis if NSCLCs are poorly differentiated and not clearly differentiated by routine light microscopy8,9,11,12.
Thyroid transcription factor-1 (TTF-1) is the strongest diagnostic marker for adenocarcinoma, while p63 is a reliable marker for squamous cell carcinoma. Currently, TTF-1 and p63 are the two principle markers capable of subtyping NSCLC8,9,11,12. Mucin stain may also be useful for identifying adenocarcinoma, likewise cytokeratin 5/6 (CK5/6) can be valuable for distinguishing squamous cell carcinoma. Napsin A is a highly specific marker for lung adenocarcinoma and p40 is an antibody that recognizes a specific p63 isoform, and is superior to p63 in specificity, with potential to replace p63 as a marker of squamous cell carcinoma8,9,11,12. In general, immunohistochemical panels containing TTF-1, p63, CK5/6, mucin stain or TTF-1, p63, CK5/6, and Napsin A are used to subtype poorly differentiated NSCLC19,20. Again, the IHC tests for histologic subtype should be minimal to preserve as much tissue as possible for molecular analysis from small biopsy and cytology samples8-11,18.
The specimen purity, defined by the ratio of malignant cells to "contaminating" non-neoplastic cells and necrotic tissue, is crucial for the sensitivity of all molecular methods; therefore, tumor cell enrichment is necessary for molecular tests, such as direct sequencing, which has relatively low sensitivity10,11,14. This can be performed by demarcating the area of interest in a paraffin block or glass slide and manual macro/microdissection. Laser capture microdissection is rarely necessary in histological specimens, but is sometimes useful for cytological specimens10,11.
DNA quality is primarily influenced by specimen fixation time, the fixation solution used, and the degree of tissue necrosis. Fixation times of 6 to 48 hours are recommended for adequate results. Formalin is the most widely used fixative, although alcohol fixed specimens show comparable results10-12.
If molecular analyses are performed sequentially, results should be available for EGFR and KRAS mutation analysis within 5 working days, ALK IHC within 2 working days, and ALK fluorescence in situ hybridization (FISH) test within 3 working days. The total turnaround time (TAT) for EGFR and ALK testing should not exceed 2 weeks (10 working days)10,11.
In practice, the duration of the order-to-report time may be delayed by several factors. The communication between clinicians and pathologists in hospital is important and poor collaboration may lead to delayed results. If the hospital is not set up to perform molecular analyses, samples are transported to specialized molecular pathology laboratories and this can also increase the order-to-report time10,11.
EGFR mutation status is the best predictor of response to EGFR-TKI therapy in NSCLC5,6,21. It is important to determine which patients should be tested for EGFR mutations, and which tests should be performed10. The most common EGFR mutations are deletions in exon 19 and the L858R point mutation in exon 21; together these comprise up to 90% of all activating EGFR mutations10,22. However, clinical EGFR mutation testing should be able to detect all individual mutations that have been reported with a frequency of at least 1% in EGFR-mutated lung adenocarcinomas10. EGFR mutations are significantly more frequent in never smokers, women, East Asians, and adenocarcinomas, but also occur in smokers, men, and Western; therefore, these clinical characteristics are not absolute markers indicating that EGFR mutation analysis is necessary10,11.
In adenocarcinoma, the prevalence of EGFR mutation is 45% in Pacific Asians and 24% in Caucasians10. A recent Korean nationwide survey reported that the overall EGFR mutation rate was 34.3% in NSCLC and 43.3% in the adenocarcinoma histologic type23.
EGFR mutations were significantly associated with adenocarcinoma in situ, minimally invasive adenocarcinoma, and lepidic- and papillary-predominant adenocarcinoma subtypes, whereas they were rarely detected in mucinous subtype tumors8,10,24. EGFR mutations have also been found in large cell carcinomas, while the mutation rate in squamous cell carcinoma is generally very low11. Therefore, EGFR mutation testing is recommended for adenocarcinoma, large cell carcinoma, NSCLC-NOS, and squamous cell carcinomas of never smokers and where only small biopsy specimens are available4.
Direct sequencing is widely used for EGFR mutation detection and remains the gold standard for gene mutation analysis, although it has low sensitivity. The method requires that at least 50% of assayed cells are malignant, corresponding to mutations in approximately 25% of the total DNA in the sample10,11. In addition to direct sequencing, various other techniques are used to detect EGFR mutations, such as pyrosequencing, Scorpion amplification refractory mutation system, and peptide nucleic acid polymerase chain reaction clamping. These new techniques are more sensitive and rapid than direct sequencing (Table 1)10,11,25,26.
Recently, EGFR mutation-specific antibodies, recognizing the two most common mutations, deletion E746-A750 in exon 19 and the L858R point mutation in exon 21, have been developed and can be used in immunohistochemical staining to identify the expressed mutated EGFR proteins. This technique is fast, inexpensive, and easily available in most pathology laboratories, although it is lacks sufficient sensitivity to be used as standard method. Although EGFR mutation-specific IHC is not currently recommended to select candidates for EGFR-TKI therapy, the method could be useful for mutation screening and as a predictive marker for patient management in the future10,11,25,27.
In practice, adequate tissue samples for diagnosis are not obtained for all patients with lung cancer. Advanced stage NSCLC is often diagnosed solely using cytology samples.
Cytology samples, such as malignant pleural effusion and fine needle aspirate, are also suitable for EGFR mutation testing. Use of cell blocks is preferred to smear preparations, due to the high tumor cell content and preservation of the original specimen. Use of more sensitive mutation testing methods is required when cytology samples with low tumor content are used10,25.
The ALK gene rearrangement is a recently identified therapeutic target for tyrosine kinase treatment in NSCLC28. The ALK gene translocation was originally identified in a subset of anaplastic large cell lymphomas. Recently, it was also discovered in NSCLC and the most common ALK fusion oncogene partner is the echinoderm microtubule-associated protein-like 4 (EML4)10,29.
EML4-ALK rearrangement is present in about 4% of all cases of NSCLC30. It is more frequent in younger patients, never/light smokers, and adenocarcinoma (predominantly solid pattern and signet ring cell subtypes)30,31. In one study, which focused on patients with adenocarcinoma histology, the frequency of EML4-ALK was 22% within the group of never or light smokers and 33% among never or light smokers without EGFR mutations32. This mutation is mutually exclusive with EGFR and KRAS mutations, with very rare exceptions. Therefore, the guidelines recommend a stepwise-testing algorithm: first, analyses of the most common mutations, EGFR and KRAS, are performed and, if the results of these are negative, testing for ALK rearrangement is undertaken10,11,31.
Currently, FISH, which has been used in clinical trials with crizotinib, is considered the standard technique for detection of ALK rearrangement and is also the Food and Drug Administration (FDA)-approved method10,11. An ALK FISH assay, using dual color break-apart probes, is commonly used. With this assay, two FISH probes of different colors (red and green) flank the common breakpoint within ALK, separating when a rearrangement occurs. ALK rearrangement appears as distinct and separated red and green signals, while non-rearrangement is seen as a fused yellow signal10,30,33. FISH allows the use of archival fresh frozen paraffin embedded tissue and can detect any fusion partner. However, the technique requires specialized equipment and expertise, is expensive, and has a relatively long TAT10,31,33.
IHC and reverse transcriptase-polymerase chain reaction (RT-PCR) are other diagnostic methods for ALK rearrangement. IHC is technically easy, relatively cheap, rapid, and considered as a useful screening method to select candidates for treatment as the ALK FISH assay10,31,33-36. RT-PCR is also technically easy, rapid, and sensitive; however, this method is only applicable for known fusion partners and it is difficult to obtain high-quality RNA in clinical practice (Table 2)10,31,33,34.
Figure 1 shows a diagnostic algorithm for histologic subtyping and molecular analysis in NSCLC diagnosis.
Recently, the development of new therapeutic agents has led accurate histologic subtyping and molecular predictive testing to be mandatory in NSCLC. In this new diagnostic approach, close communication between pulmonologist, radiologist, pathologist, and oncologist, involved in tissue acquisition, pathology diagnosis, molecular analysis, and treatment decisions is necessary. Specifically, each hospital should have a multidisciplinary strategy to obtain adequate tissue samples, process them in a pathology laboratory, classify them into histologic subtypes, and preserve materials for molecular testing, to ensure fast diagnosis without delay.
Ultimately, we expect therapeutic outcomes in NSCLC to improve with personalized therapy, which is driven by the selection of appropriate treatments based on histologic subtype and molecular status, including EGFR mutations and ALK rearrangement.
References
1. Scagliotti GV, Parikh P, von Pawel J, Biesma B, Vansteenkiste J, Manegold C, et al. Phase III study comparing cisplatin plus gemcitabine with cisplatin plus pemetrexed in chemotherapy-naive patients with advanced-stage non-small-cell lung cancer. J Clin Oncol 2008;26:3543-3551. PMID: 18506025.


2. Scagliotti G, Brodowicz T, Shepherd FA, Zielinski C, Vansteenkiste J, Manegold C, et al. Treatment-by-histology interaction analyses in three phase III trials show superiority of pemetrexed in nonsquamous non-small cell lung cancer. J Thorac Oncol 2011;6:64-70. PMID: 21119545.


3. Sandler A, Gray R, Perry MC, Brahmer J, Schiller JH, Dowlati A, et al. Paclitaxel-carboplatin alone or with bevacizumab for non-small-cell lung cancer. N Engl J Med 2006;355:2542-2550. PMID: 17167137.


4. National Comprehensive Cancer Network (NCCN). NCCN clinical practice guidelines in oncology. Non-small cell lung cancer version 2.2013 [Internet]. Fort Washington: National Comprehensive Cancer Network; 2013. cited 2013 Aug 19. Available from: http://www.nccn.org/professionals/physician_gls/f_guidelines.asp#nscl.
5. Mok TS, Wu YL, Thongprasert S, Yang CH, Chu DT, Saijo N, et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N Engl J Med 2009;361:947-957. PMID: 19692680.


6. Zhou C, Wu YL, Chen G, Feng J, Liu XQ, Wang C, et al. Erlotinib versus chemotherapy as first-line treatment for patients with advanced EGFR mutation-positive non-small-cell lung cancer (OPTIMAL, CTONG-0802): a multicentre, open-label, randomised, phase 3 study. Lancet Oncol 2011;12:735-742. PMID: 21783417.


7. Kwak EL, Bang YJ, Camidge DR, Shaw AT, Solomon B, Maki RG, et al. Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N Engl J Med 2010;363:1693-1703. PMID: 20979469.



8. Travis WD, Brambilla E, Noguchi M, Nicholson AG, Geisinger KR, Yatabe Y, et al. International Association for the Study of Lung Cancer/American Thoracic Society/European Respiratory Society international multidisciplinary classification of lung adenocarcinoma. J Thorac Oncol 2011;6:244-285. PMID: 21252716.



9. Travis WD, Brambilla E, Noguchi M, Nicholson AG, Geisinger K, Yatabe Y, et al. Diagnosis of lung cancer in small biopsies and cytology: implications of the 2011 International Association for the Study of Lung Cancer/American Thoracic Society/ European Respiratory Society classification. Arch Pathol Lab Med 2013;137:668-684. PMID: 22970842.


10. Lindeman NI, Cagle PT, Beasley MB, Chitale DA, Dacic S, Giaccone G, et al. Molecular testing guideline for selection of lung cancer patients for EGFR and ALK tyrosine kinase inhibitors: guideline from the College of American Pathologists, International Association for the Study of Lung Cancer, and Association for Molecular Pathology. J Thorac Oncol 2013;8:823-859. PMID: 23552377.



11. Thunnissen E, Kerr KM, Herth FJ, Lantuejoul S, Papotti M, Rintoul RC, et al. The challenge of NSCLC diagnosis and predictive analysis on small samples: practical approach of a working group. Lung Cancer 2012;76:1-18. PMID: 22138001.


12. Moreira AL, Thornton RH. Personalized medicine for non-small-cell lung cancer: implications of recent advances in tissue acquisition for molecular and histologic testing. Clin Lung Cancer 2012;13:334-339. PMID: 22424871.


13. Rivera MP, Mehta AC. American College of Chest Physicians. Initial diagnosis of lung cancer: ACCP evidence-based clinical practice guidelines (2nd edition). Chest 2007;132(3 Suppl):131S-148S. PMID: 17873165.


14. Kulesza P, Ramchandran K, Patel JD. Emerging concepts in the pathology and molecular biology of advanced non-small cell lung cancer. Am J Clin Pathol 2011;136:228-238. PMID: 21757595.


15. Alsharif M, Carlo-Demovich J, Massey C, Madory JE, Lewin D, Medina AM, et al. Telecytopathology for immediate evaluation of fine-needle aspiration specimens. Cancer Cytopathol 2010;118:119-126. PMID: 20544707.


16. Navani N, Brown JM, Nankivell M, Woolhouse I, Harrison RN, Jeebun V, et al. Suitability of endobronchial ultrasound-guided transbronchial needle aspiration specimens for subtyping and genotyping of non-small cell lung cancer: a multicenter study of 774 patients. Am J Respir Crit Care Med 2012;185:1316-1322. PMID: 22505743.



17. Sakairi Y, Nakajima T, Yasufuku K, Ikebe D, Kageyama H, Soda M, et al. EML4-ALK fusion gene assessment using metastatic lymph node samples obtained by endobronchial ultrasound-guided transbronchial needle aspiration. Clin Cancer Res 2010;16:4938-4945. PMID: 20926401.


18. Aisner DL, Marshall CB. Molecular pathology of non-small cell lung cancer: a practical guide. Am J Clin Pathol 2012;138:332-346. PMID: 22912349.


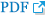
19. Loo PS, Thomas SC, Nicolson MC, Fyfe MN, Kerr KM. Subtyping of undifferentiated non-small cell carcinomas in bronchial biopsy specimens. J Thorac Oncol 2010;5:442-447. PMID: 20195168.


20. Mukhopadhyay S, Katzenstein AL. Subclassification of non-small cell lung carcinomas lacking morphologic differentiation on biopsy specimens: utility of an immunohistochemical panel containing TTF-1, napsin A, p63, and CK5/6. Am J Surg Pathol 2011;35:15-25. PMID: 21164283.


21. Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW, et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med 2004;350:2129-2139. PMID: 15118073.


22. Sharma SV, Bell DW, Settleman J, Haber DA. Epidermal growth factor receptor mutations in lung cancer. Nat Rev Cancer 2007;7:169-181. PMID: 17318210.


23. Kim WS, Choi YD, Seo JW, Han JH, Kim MJ, Kim L, et al. Epidermal growth factor receptor mutation analysis of non-small cell lung cancer in South Korea: summary from a nationwide survey. J Lung Cancer 2010;9(Suppl 2):S38.
24. Yoshizawa A, Sumiyoshi S, Sonobe M, Kobayashi M, Fujimoto M, Kawakami F, et al. Validation of the IASLC/ATS/ERS lung adenocarcinoma classification for prognosis and association with EGFR and KRAS gene mutations: analysis of 440 Japanese patients. J Thorac Oncol 2013;8:52-61. PMID: 23242438.


25. Ellison G, Zhu G, Moulis A, Dearden S, Speake G, McCormack R. EGFR mutation testing in lung cancer: a review of available methods and their use for analysis of tumour tissue and cytology samples. J Clin Pathol 2013;66:79-89. PMID: 23172555.


26. Pao W, Ladanyi M. Epidermal growth factor receptor mutation testing in lung cancer: searching for the ideal method. Clin Cancer Res 2007;13:4954-4955. PMID: 17785543.


27. Kawahara A, Taira T, Azuma K, Tominaga M, Hattori S, Kawahara M, et al. A diagnostic algorithm using EGFR mutation-specific antibodies for rapid response EGFR-TKI treatment in patients with non-small cell lung cancer. Lung Cancer 2012;78:39-44. PMID: 22858448.


28. Soda M, Choi YL, Enomoto M, Takada S, Yamashita Y, Ishikawa S, et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature 2007;448:561-566. PMID: 17625570.


29. Rikova K, Guo A, Zeng Q, Possemato A, Yu J, Haack H, et al. Global survey of phosphotyrosine signaling identifies oncogenic kinases in lung cancer. Cell 2007;131:1190-1203. PMID: 18083107.


30. Solomon B, Varella-Garcia M, Camidge DR. ALK gene rearrangements: a new therapeutic target in a molecularly defined subset of non-small cell lung cancer. J Thorac Oncol 2009;4:1450-1454. PMID: 20009909.


31. Murakami Y, Mitsudomi T, Yatabe Y. A screening method for the ALK fusion gene in NSCLC. Front Oncol 2012;2:24PMID: 22655265.



32. Shaw AT, Yeap BY, Mino-Kenudson M, Digumarthy SR, Costa DB, Heist RS, et al. Clinical features and outcome of patients with non-small-cell lung cancer who harbor EML4-ALK. J Clin Oncol 2009;27:4247-4253. PMID: 19667264.



33. Weickhardt AJ, Aisner DL, Franklin WA, Varella-Garcia M, Doebele RC, Camidge DR. Diagnostic assays for identification of anaplastic lymphoma kinase-positive non-small cell lung cancer. Cancer 2013;119:1467-1477. PMID: 23280244.


34. Marchetti A, Ardizzoni A, Papotti M, Crino L, Rossi G, Gridelli C, et al. Recommendations for the analysis of ALK gene rearrangements in non-small-cell lung cancer: a consensus of the Italian Association of Medical Oncology and the Italian Society of Pathology and Cytopathology. J Thorac Oncol 2013;8:352-358. PMID: 23407559.


35. Paik JH, Choe G, Kim H, Choe JY, Lee HJ, Lee CT, et al. Screening of anaplastic lymphoma kinase rearrangement by immunohistochemistry in non-small cell lung cancer: correlation with fluorescence in situ hybridization. J Thorac Oncol 2011;6:466-472. PMID: 21258247.


36. Paik JH, Choi CM, Kim H, Jang SJ, Choe G, Kim DK, et al. Clinicopathologic implication of ALK rearrangement in surgically resected lung cancer: a proposal of diagnostic algorithm for ALK-rearranged adenocarcinoma. Lung Cancer 2012;76:403-409. PMID: 22129856.


Figure 1
A diagnostic algorithm for histologic subtyping and molecular analysis in non-small cell lung cancer (NSCLC) diagnosis. SCLC: small cell lung cancer; H&E: hematoxylin and eosin; SQCC: squamous cell carcinoma; ADC: adenocarcinoma; NOS: not otherwise specified; TTF-1: thyroid transcription factor-1; CK: cytokeratin; IHC: immunohistochemistry; EGFR: epidermal growth factor receptor; TKI: thyrosine kinase inhibitor; ALK: anaplastic lymphoma kinase; FISH: fluorescence in situ hybridization.
Table 1
Methods for EGFR mutations testing in NSCLC
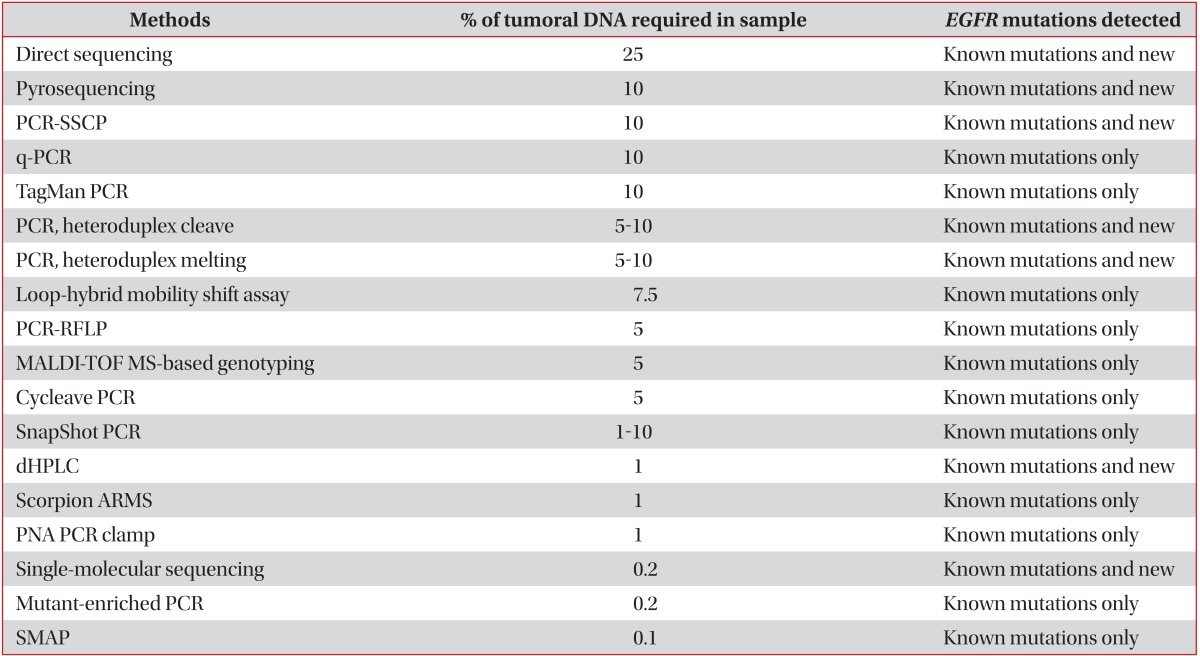
EGFR: epidermal growth factor receptor; NSCLC: non-small cell lung cancer; PCR-SSCP: polymerase chain reaction (PCR) plus single-strand conformation polymorphism; q-PCR: quantitative PCR; PCR-RFLP, PCR-restriction fragment length polymorphism; MALDI-TOF MS: matrix-assisted laser desorption/ionization time-of-flight mass spectrometry; dHPLC: denaturing high performance liquid chromatography; ARMS: amplified refractory mutation system; PNA PCR clamp: peptide nucleic acid PCR clamp; SMAP: smart amplification process.



 PDF Links
PDF Links PubReader
PubReader Full text via DOI
Full text via DOI Print
Print Download Citation
Download Citation